CARTANA in situ sequencing was just bought by 10X Genomics for $41.2 million. They provide a complement to the Visium technology that generates beautiful gene expression maps of tissue sections as seen in Fig 1c of a recent bioRxiv report (see below).

The method was also used in a recent Cell publication from the VIB-KU Leuven Center for Brain and Disease Research “Spatial Transcriptomics and In Situ Sequencing to Study Alzheimer’s Disease”. They applied the Cartana technology to Alzheimer’s disease development and progression to confirm visium gene expression results.
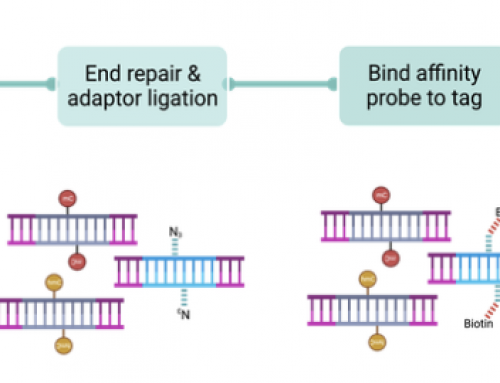
How it works: analyse either fresh/fixed frozen or FFPE samples and rapidly create single-cell gene expression maps of up to hundreds of genes. Padlock probes hybridise to target-specific RNAs. The closed circle is then copied locally by a rolling circle amplification (RCA) to generate a “spot” of the transcript that includes an in situ barcode. Barcodes are decoded by multiple cycles of probe hybridisation to generate highly-informative maps of gene expression for the targeted genes while preserving tissue morphology.
The technology was originally published in Nature Methods as pciSeq (probabilistic cell typing by in situ sequencing) by Mats Nilsonn’s group. In the paper they also discuss the pros and cons of the various methods for multiplexed in situ RNA detection and cell calling (well worth a read).







Leave A Comment