I recently had a days training at Oxford Nanopore’s labs on their new cDNA-Seq kit and ran the resulting libraries on the GridION (one of the first customers to do so). A lot of people are watching RNA-Seq developments at ONT very closely. The London Calling session on transcriptome analysis included presentation on cDNA-Sequencing work, and although arguably the most exciting work presented was on direct RNA-Seq, I think many people will get their first taste of RNA-Seq on Nanopores via one of the two cDNA-Seq kits (PCR-free or PCR+) being released today – the Tweet below from @Genomique_ENS was the first notification this was coming.
New cDNA strand switching protocol by @nanopore tested @Genomique_ENS. Almost 10 million 1D reads ! Where will it stop ????#nanoporeconf
— Génomique ENS (@Genomique_ENS) May 4, 2017
ONT are offering the first easily accessible platform for full-length cDNA sequencing (see comparisons to ISO-Seq later in this post), and are generating up to 10 million reads per flowcell. Their new kit offers a window into the transcriptome we’ve simply never had – an easy method to sequence full-length mRNA transcripts.
At perhaps as little as £200 per sample it compares well to Illumina sequencing…how long will it be before we leave behind short reads for RNA-Seq?
Read on for more about this new kit from me; or follow these links for more information from ONT: Some videos, their white paper (needs registration), an RNA starter pack on the store, or other kits for people with a MinION (and a useful comparison table)
New today! Direct or PCR-based cDNA analysis kits for @nanopore. Or go for direct RNA analysis #RealRNASeq https://t.co/XiaHgEou26 pic.twitter.com/ZL9Naqasyr
— Oxford Nanopore (@nanopore) June 26, 2017
Why use full-length cDNA sequencing:
Northern blotting, RTqPCR, SAGE, Microarrays, RNA-Seq, cDNA-seq, Direct RNA-Seq; the evolution of methods for RNA analysis has led to many exciting discoveries along the way. That RNA splicing occurs is well known. It occurs in perhaps as much as 95% of genes, with an average of more than five isoforms per gene. The development of exon and tiling arrays by Affymetrix hinted at the complexity of the transcriptome, but it was RNA-Seq that allowed us to really dig into the transcriptome.
However most of the published RNA-Seq data is from Illumina technology where short reads (single-end 50bp or paired-end 75bp) from short cDNAs (average 150-200bp) mean that isoforms must be detected computationally for almost all Human genes.
Full-length cDNA-sequencing (and direct RNA in the future) allows researchers to detect isoforms perhaps as easily as counting genes in traditional RNA-Seq. Simply reverse-transcribing and sequencing seems like a simple enough approach – but I’m sure there will be lots of challenges that we’ll need to resolve, in the lab and computationally.
The recent BioRxiv report from Mark Akeson & Christopher Vollmers’ labs at UCSC showed how even single-cell transcriptomes of mouse immune cells could be analysed by sequencing full-length cDNA molecules on MinION. Sounds like the future is here already!
Comparisons to RNA-Seq on Illumina:
Illumina sequencing of stranded mRNA or Total RNA-seq libraries has been very widely adopted and consequently “RNA-Seq” is the most widely published NGS method. Manolis Dermitzakis lab, in one of two papers from 2010 that really kick-started large-scale RNA-Seq analysis, demonstrated how just 10 million sequencing reads could give the same dynamic range as microarrays with better quantification of alternative and highly abundant transcripts. The ENCODE Guidelines v3 generally call for 20-30 million reads per sample. However higher numbers of longer reads are often used for whole transcriptome analysis – maybe 50-100M paired-end 75bp or even 150bp reads.
My own lab makes ~3000 RNA-Seq libraries a year. We are usually performing differential gene expression analysis and use stranded mRNA kits from Illumina to generate 10-20 million SE50bp reads per sample. This is a highly effective way to do DGE. It is sensitive, pretty specific, and cheap at just £150 per sample.
For me the biggest unanswered question is how do you compare Illumina and ONT RNA-Seq reads? Are nanopore reads worth more? Will smaller numbers of reads give the same sensitivity/specificity for DGE, and DIE? How will an Illumina short-read RNA-Seq data compare to a Nanopore full-length cDNA-Seq data? This is going to depend very much on the questions being asked by the biologist.
In the Byrne et al BioRxiv report they state “even with the relative low number of reads produced, ONT RNA-Seq gene-expression quantification largely detects the same genes as Illumina RNA-Seq” .
Long is obviously better for isoform analysis and 1 ONT read of 2000bp could equal 40, 50 even 100s of Illumina SE50bp reads. But counting requires molecules so I suspect we’ll still want millions of nanopore reads per sample. The MinION is capable of generating 10M reads when fed full-length cDNA, but if fragmented to a tenth of that length should generate close to 100M reads. However until ONT demonstrate read levels this high ONT remains relatively costly – for differential gene expression!
However for isoform analysis ONTs cDNA-Seq protocol is likely to be very attractive and I suspect likely to be run alongside Illumina RNA-Seq to generate combination datasets. This would be a similar approach to that being used for genome sequencing where a mix of Illumina and, PacBio, 10X Genomics, BioNano, or MinION data generates the best assemblies.
I’ve left a cost comparison to a future post as I’m anticipating lots of interesting discussion around this topic over the next few weeks and months!
Comparisons to ISO-Seq:
Iso-Seq from Pacific Biosciences is capable of generating full-length reads spanning transcript isoforms from polyA-tail to 5′ end. The system has been reasonably well adopted and there are many publications using it to investigate isoform abundance. One of the early papers: A single-molecule long-read survey of the human transcriptome from Michael Snyder’s Stanford lab reported sequencing the transcriptome with circular consensus reads generating 2-fold coverage of full-length cDNAs ranging from 1.5-2.5Kb; including forward and reverse cDNA strands. When the authors compared their results to GENCODE transcriptome annotation they found about 10% of their data were novel unannotated transcripts.
Iso-Seq requires cDNA to be size fractionated for sequencing to reduce bias coming from fragment length. This is a headache for users but it remains to be seen if the ONT cDNA protocol is unbiased, or whether it will require a similar approach.
According to PacBio it is now possible to generate reads from cDNAs of up to 10kb in length. The most recent report on BioRxiv from Workman et al generated 8.6Gb of sequencing data, or 2.6M reads, from four size fractions (1-2kb, 2-3kb, 3-6kb, 5-10kb) on 40 SMRT Cells. The longest read was of acetyl-coA carboxylase 1 spanning the 7kb length of its coding sequence.
Our first experiments have generated 4.5M, 5.2M and 6.5M cDNA reads on the MinION which aces the PacBio SMRT cells. We’ve lots to do to be able to say much more than “we got more reads” but I’m excited to get the analysis underway.
The protocol:
Developed by Phillip James in the applications group at ONT, the cDNA-Seq kit allows users to prepare libraries in about 5 hours ready for MinION, GridION or PromethION sequencing.
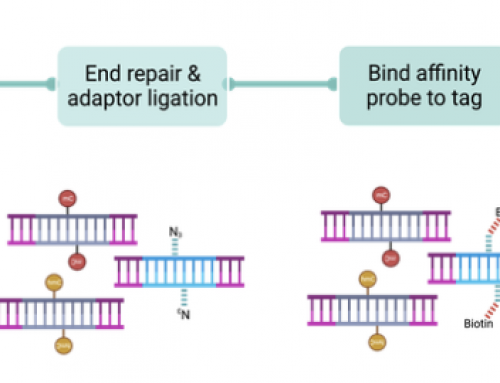
The protocol uses a cDNA strand switching approach, which enriches for full-length cDNA molecules where reverse transcription has reached the 5’ end of the mRNA. Fragmented mRNAs will also be enriched so RNA quality is critical – as will be methods to remove fragmentation products from Differential Isoform Expression (DIE-Seq) analysis.
RNA input recommendations are for 50 ng poly A tailed mRNA, or 500ng of Total RNA. In future tests of this technology we’ll be looking at the impact of RNA input to the kit as we want to maximise full-length transcripts so don’t want to overload the RT reaction. However I’d much prefer an assay that works robustly from Total RNA rather than quantified mRNA. It may be possible to simply add an oligo-dT mRNA pull-down in the first step and to use the pull down oligo-dT as the cDNA synthesis primer.
• Anneal cDNA synthesis primer: mix RNA with cDNA synthesis primer, dNTPs and water. Mix by flicking, spin, heat to 65 °C for 5 min. Snap cool on ice.
• Add strand-switching reagents: prepare a master of SuperScript IV, RNase OUT, DTT, and strand switching primer. Add to the snap cooled, annealed mRNA. Heat at 50 °C for 10 min, 42 °C for 10 min, 80 °C for 10 min, and cool and hold at 4 °C ready for cDNA PCR.
• Enrich full-length cDNAs by PCR: Mix 2 x LongAmp Taq Mastermix, cPRM primers and cDNA. PCR for 11-18 cycles 95 °C for 15 sec, 62 °C for 15 sec, 65 °C for 50 second per Kb; with a final 65 °C for 6 min, then cool and hold at 4 °C ready for clean-up.
• Ampure XP clean-up: Add 0.8x SPRI (40ul to a 50ul PCR), wash twice with 70% ethanol, resuspend after drying in elution buffer. This is your pre-adapted library.
• Library prep: Quantify your library using Qubit HS DNA quant kit. You should get 150-700ng; if not reduce/increase PCR cycles as required. Add cAMX to ~500ng of library. Gently mix on Hula mixer for 5 min at room temp. This is your adapted library.
• Ampure XP clean-up: Add 0.8x SPRI (20ul to a 25ul adapted library), wash twice with the buffer supplied by ONT (not ethanol), resuspend. This is your sequencing library.
• Prep and wash your flowcell.
• Load the library and start sequencing!
Our initial results: We sequenced the MAQC RNAs (Ambion Brain and Stratagene UHRR), as these are very well characterised already, and generated 4.5M, 5.2M and 6.5M reads on MinION; and are awaiting download of our GridION data. We will compare differential gene expression results to those obtained on Illumina with short reads to try and determine how many Illumina reads one MinION read equates to.
cDNA primer design: as the MinION sequences full-length cDNA molecules it would make a lot of sense to add unique molecular identifiers to the cDNA synthesis primer. This would be a very simple modification but may prove useful in reducing the impact of amplification bias on analysis as duplicate molecules would be removed allowing only unique cDNA molecules to go forward to differential isoform analysis.
Nanopore sample handling tips and tricks: “Mix by flicking” is going to be the standard method for mixing anything in a long-read method. Pipetting, especially small volumes with narrow bore tips, is simply too rough and will break template strands and cDNA molecules leading to lower numbers of full-length cDNA reads. Be gentle!
If you’ve thoughts about RNA-Seq/cDNA-Seq on ONT and its comparison to Illumina please leave a comment of post on Twitter.






IsoSeq on the Sequel makes size fraction in optional. The Sequel has a much reduced library molecule length bias.
Thanks for your comments Lutz, I’ve not looked at IsoSeq recently enough. Thanks for the prompt.