Index-swapping appears to be driven by excess adapter/primer (Illumina whitepaper). The take-home messages are 1) to use UDIs (unique dual-indexes), and 2) clean-up your libraries to remove left-over adapter/primer. Later in this post I’m going to work through one solution for getting rid of any remaining adapter/primer in an NGS library; but first I’m going to talk about reducing the impact of index-swapping on RNA-Seq experiments.
RNA-Seq and index-swapping
For many projects index-swapping in RNA-Seq is unlikely to be a big problem. Since sample groups are generally replicated (biological replicates please) the random nature of index-swapping means a slight increase in background noise that is likely to make low-expression transcripts slightly harder to detect – but many users are not primarily focused on low-expressors anyway.
However for any RNA-Seq experiment it is well worth performing standard QCs on the RNA and on the final libraries before sequencing.
3 simple steps to avoid index-swapping problems in RNA-Seq
- Check RNA quality and quantity before starting
- Be meticulous about your bead cleanups (read about how SPRI-beads work here)
- Check library size, quality and quantity before pooling
We’ve just purchased and Agilent TapeStation for high-throughput QC so we can check every library for size and adapter contamination. Running 96 RNA-Seq libraries on the BioAnalyser was such a pain we had defaulted to checking just 12 at random. This was fine for verifying that our RNA-Seq preps were working, but limiting in that a low-quality library could be missed and included in the run. This would be particularly bad for a sample that had failed as there would most likely be lots of adpater/PCR-primer left over after prep. This simple QC run on BioAnalyser/TapeStation should give you a good idea about how much adapter/primer remains in your library; basically if there is anything detectable then do another clean-up!
Size-selection of library pools to remove left-over adapter/primer
We’ve been using SPRI-bead cleanups for many years and they generally do a good job of removing the smaller adapter/primer molecules from a library prep…bead cleanups are great! However for some library types this cleanup does not work quite so well.
Libraries that may be more badly affected might be smallRNA as the difference between insert size and adapter dimer is only tens of bases (see figure below). Even a standard RNA-Seq library, usually around 270bp, might be less affected by index-swapping, and better for RNA-Seq analysis, by increasing its size to 350bp. This would result in easier size selection during cleanup and allow for PE150 sequencing – with less wasted sequence from overlaps in the reads.

For even finer control over size-selection a gel purification of the final library might be the best option. This is what is recommended in the samllRNA protocols anyway. But for just about any library this would most likely remove almost all excess adapter/primer. With instruments like the Pippin Prep this would be pretty simple. This instrument is recommended by Illumina for some workflows already, it allows users to select narrow fragment distributions, reproducibly across experiments with minimal effort.
Removing excess adapter/primer enzymatically
ExoSAP: Anyone who did lots of Sanger sequencing is likely to have come across ExoSAP/ExoSAP-IT. This is a cocktail of Exonuclease and Shrimp Alkaline Phosphatase used for enzymatic cleanup of amplified PCR products. The Exonuclease I degrades the left-over PCR primers and the Shrimp Alkaline Phosphatase dephosphorylates any left-over dNTPs. Since ExoSAP can be simply added after the final PCR it would be an easy solution to cleanup NGS libraries (see NEBs handy guide).

I’ve never seen this method proposed in an NGS pipeline. It is something easily tested, but with ehe “busyness” of most labs is an experiment that simply gets left on the back-burner. I might just look at testing this in my lab this year though! A word of caution, Exonuclease1 was reported to nick DNA in this 1979 paper, this might still be a problem (As pointed out by Chris in his comment below it was not the Exo1 causing the nicking!).
Reducing contamination in the lab
While thinking about the application of ExoSAP to NGS I remembered an idea we’d discussed many years ago to help reduce the chances of library contamination during prep in the lab. One area of concern is that PCR amplified libraries can contaminate the next set of templates, or the next library-prep process.
A UNG step is very commonly used is in qPCR assays to prevent PCR carryover contamination, and reduce false positives. It works by degrading PCR products from previous PCR amplifications without degrading native nucleic acid templates. The same process could be applied to NGS library prep and reduce contamination between preps. AmpErase Uracil N-Glycosylase (UNG) excises Uracils incorporated into PCR products – in our case NGS libraries – in place of Thymines. Native DNA, or pre-PCR adapter-ligated NGS libraries are not affected, but any contaminating PCR-amplified, and Uracil containing, NGS library molecules are degraded by the UNG step.
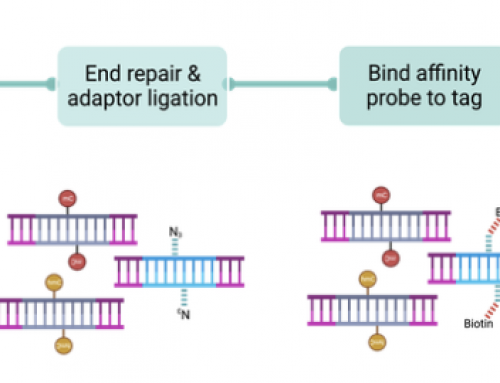
If Uracil containing indexed adapters are used but PCR is completed with universal primers (P5 & P7 only) then conceivably index-swapping could also be reduced by using UNG to degrade left over adapters instead, but not the PCR amplified templates (obviously neither method is not going to work for PCR-free libraries).







Looking quickly at the Rosamond et al 1979 paper cited here, I think perhaps there’s a been a misinterpretation — the nonspecific nicking they’re referring to is a function of the RecBC enzyme, not Exonuclease I.
Thanks for pointing this out Chris, I’ve updated the post to reflect your comments.
Of course the UNG method won’t work for dUTP stranded RNA-seq libraries, where one strand is marked with uracil. You would need to use an enzyme that extends through uracil, meaning you’d lose strand specificity.