Illumina released a white paper this week describing their efforts to understand the index-swapping issues with the ExAmp chemistry. This story has been covered many times and got lots of Tweets in the days since the BioRxiv paper from Stanford was uploaded. Megan Molteni at Wired wrote a lengthy piece that included comments from Omead Ostadan (EVP at Illumina). Megan reported that Illumina “has known about barcode hopping (index-swapping) for 10 years” and that after Stanford (Sinha et al) complained that “We realized we had to move quickly to characterize the problem.”
The white paper was most likely in preparation at Illumina when the BioRxiv paper was uploaded. I’ve been talking to Illumina for almost a year about indexing issues including index-swapping, and clearly they’ve taken Rahul’s work seriously. Illumina have been working on understanding the root cause and identifying mitigations we can easily roll out in our labs, and describe much of this work in the white paper. However I’m sure it will have been rushed out ASAP after the BioRxiv paper landed on Francis deSouza’s desk.
Megan’s piece is also the first report of financial impact due to of this issue the Weissman lab said it lost $1 million (salaries + consumables etc) and is quoted as saying he “wishes someone would declare a state of emergency”.
More about Illumina’s white paper in a sec first I wanted to point readers to some other coverage of this issue…but first wanted to summarise what I think users need to know:
- Index-swapping affects most library types – but at a very low level
- Users working on low-frequency variant detection should think most carefully about the impact
- Most users will be unaffected with clean libraries i.e. no adapter/primer contamination
Ethan Linck at The Molecular Ecologist writes about the BioRxiv paper…pointing to Gavin Sherlock’s marvellous play-on-words Tweet:
“I think this is a genuine cluster fuck.”
Ethan highlights the kinds of experiments that may be adversely affected and points out that, as a preprint, we need to see if this makes into realprint after peer review. Ethan also says we need to see “if other labs can replicate their results” I am sure they will and more and more reports will come out.
GenomeWeb have just covered this story and summarise the information. They interviewed Rahul Sinhar at Stanford and also quote Rasmus Nielsen (University of California, Berkeley) as saying his group has seen the same problem at similar rates of 5-10% in multi-species experiments where the problem jumped out at them.
GenomeWeb also quoted Tim DeSmet (Broad Institute) as saying they’ve been using unique dual indexes for several years, although he did say the Broad was affected by index-swapping during his AGBT presentation in Ilumina’s workshop. The Broad pipeline automatically filters reads that are index-swapped, so perhaps the Broad could comment on how widespread the issue is? And Gary Schroth (@Genomics_Guy at Illumina), confirmed to GenomWeb that “Index switching has been a known phenomenon since the early days of next-generation sequencing”.
In blog posts to UC Davis Genome Centre users Lutz Froenicke highlighted that index-swapping is most likely to affect studies looking for low abundance mutations. UCDavis first alerted their users to the issue in early April (thanks for including a link to Enseqlopedia), and most recently pointed users to Illumina white paper and attempt to reassure users that with clean sequencing libraries (no detectable free adapter/index on Bioanalyser) the majority of applications will be minimally affected. Lutz also pointed to some issues with the Sinha et al experiments that might mean it was likely to be badly affected by an issue most of us might be able to consider as noise – however I wanted to add some comments of my own (and must get in touch with Lutz/Rahul).
In his post Lutz stresses that index-swapping is an issue users can help to fix by making clean libraries, “Ugly things can happen when sequencing really ugly libraries”, that we (Illumina and users) need to investigate this more thoroughly, and that we should look at mitigations like improving magnetic bead cleanup, gel-based size selection, Exonuclease1 digestion (as suggested on SEQanswers), and using uniquely dual-indexed adapters (on Enseqlopedia of course ;-)). I’m going to write a follow-up post about the use of UNG or ExoSAP over the weekend!
- The Sinha et al study used single-cell RNA-seq SMART-Seq library preps that were “lengthy” and “not typical” . I don’t think we should focus this on the SMART-Seq protocol, all applications PCR-free or PCR+, single-cell or bulk are affected – but to differing degrees. That said plate-based PCR index addition is fraught with contamination possibilities where non-unique dual-indexes are used!
- The authors sequenced “[low quality libraries with large amounts of free primers]”. The Sinha et al bioanalyser traces do show lots of free adapter – we’d certainly suggest users clean these up – and this is likely to have been one of the two major factors contributing to the 5-10% index-swapping in their experiment.
- The authors“[sequenced libraries with an insert size much longer than recommended for HiSeq 4000 sequencing]” (mean fragment length of >800bp versus the recommended <500bp). Large molecules won’t cluster efficiently (in ExAmp or traditional clustering) meaning small molecules will be favoured by ExAmp for both clustering and index-swapping. This is the second major factor contributing to the 5-10% index-swapping in their experiment.
- That “the Stanford lab seems to work exclusively with Nextera style library preps – which often result in libraries with out-of-spec inert sizes.” Again I don’t think we should focus on one protocol. It is not the short PCR indexes that are the problem – this issue affects TruSeq PCR-free even worse than PCR+. Transposase preps are very variable and users of these library prep kits need to pay attention to insert size. We’ve seen more of an issue with wasted sequence in rapid exomes – but the 50ng input means they have a place in our library prep arsenal.

“Ugly things can happen when sequencing really ugly libraries”
Illumina’s whitepaper
This document describes how index hopping may occur, how Illumina measures it, and some best practices to mitigate its impact. They confirm the issue as molecular recombination of indexes, i.e. “Index Hopping” and that this arises primarily through contamination of libraries with excess free Adapters/Primers. Most importantly users are reassured that: 1) most experiments are likely to be unaffected by this issue, and that 2) the level of index swapping is unlikely to be higher than 1% except in specific circumstances (although we have not directly confirmed this and if you are trying to detect/quantify low-frequency variants then you should think about how you can detect this in your experiments).
The two figures reproduced below show how index swapping increases significantly with free primer/adapter. And how both PCR-free and PCR+ libraries are affected, but that HiSeq 2500 (gotta love that instrument) is much less affected (and probably for different reasons).


Illumina offer some best practices to reduce index swapping and in particular refer users to their pooling guidelines for Dual-Indexed sequencing. Whilst pooling the 8×12 PCR indexes is a pain and is probably best done on a robot, if these were re-arrayed before use pooling could be done with a multi-channel pipette pretty easily.
- Store pooled libraries at -20C, avoid multiple freeze-thaw or room-temperature storage.
- Remove as much free adapter/primer as possible and verify libraries on Bioanalyser.
- Pool libraries with Unique Dual Indexes where possible – this will allow any index-hopping to be detected, measured and removed.
In their experiments Illumina used Unique Dual Index (UDI) combinations. To make this easy for anyone running larger pooled experiments e.g. 96plex RNA-Seq we’ll need Illumina and other companies to make these indexes for us. I’m going to write a follow-up post about index design over the weekend!
A note on nomenclature
There are many terms being used to describe this issue and that will most likely make it harder for users to gather all the evidence in one place. What is clear is that indexes that you think should only appear on sample 1 on a pool also appear on sample 2 (and sample n…). How it happens is important if you’re trying to fix the root cause e.g. extension in ExAMp from free-primer is different to contamination in a lab during PCR setup, or from mis-incorporation during primer synthesis, or even from sequencing errors. But when talking about the issue I think what we care about is it’s impact.
I’m not sure which of the terms being used, Index mis-assignment, Index-hopping, or Index-swapping, is best – let me know what you think in the comments!
Summary
Illumina are working hard to fix this issue, but similarly to Rahul’s feelings, as expressed on GenomeWeb, I’m disappointed they’ve taken so long to publicly acknowledge an issue that could dramatically impact a few customers, especially since low frequency variant detection is one of the methods affected (Grail, Grail, Grail). I’ll end with his final quote from the GenomeWeb article… “I’m surprised that it took the BioRxiv paper for Illumina to come out with anything”.
Please watch out for and comment on the follow up posts coming next week on index design and the use of UNG/ExoSAP to reduce this problem to zero.


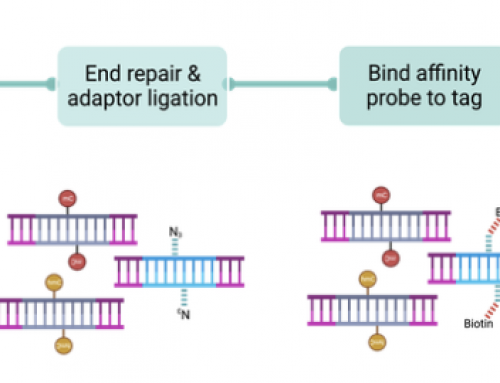

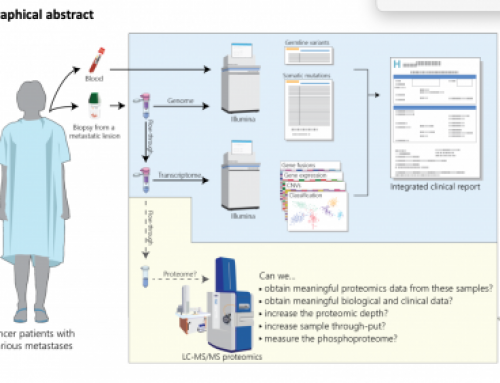




[…] downstream analysis and interpretation e.g. increased variant calling background. Incidentally, index-hopping is not a problem based on their analysis of internal […]