Introducing NovaSeq
This is the second part of my “everything you wanted to know about NovaSeq blog and in it I’ll finish up by discussing some of the challenges the instrument presents for Core Labs and their users, and the costs of running NovaSeq compared to other sequencers. At the end of this post I talk about whether you should purchase a NovaSeq or not. I’ll continue to refer to Mick’s table (thanks Mick) and the images in the gallery below.
- A comparison table of Illumina sequencers
- NovaSeq data quality %Q30
- NovaSeq data quality comparison
- NovaSeq data quality across the human genome
- NovaSeq 2colour SBS
- NovaSeq 6000 vs 5000
- NovaSeq cost per GB compared to the HiSeq family
Limitations of NovaSeq in a core-lab:
I should immediately start by pointing out that I can only talk directly about limitations my lab faces, other cores are set up differently and may have different user requirements.
My lab is a genomics core in a cancer research institute and we generate data for 11 Departments/Institutes across Cambridge. This means we run a huge variety of sequencing libraries. Whilst NovaSeq is being lauched with a much wider list of supported applications than HiSeq 4000 was it is unclear if all NGS applications will be supported (almost certainly not), or will run efficiently on NovaSeq (probably not). However we have demonstrated lots of successes in porting sequencing over to HiSeq 4000 so I would anticipate most sequencing could be completed on NovaSeq (maybe with a 2500 left in rapid mode for a few projects).
Three issues need to be resolved before we can switch the majority of sequencing over to NovaSeq; 1) pooling of multiple libraries, 2) confirmation of comparability between 4-colour and 2-colour SBS, and 3) fixing ExAmp clustering’s index swapping problems.
1) Pooling of multiple libraries: Most sequencing libraries submitted to my lab are for just one or two lanes. In 2016 we ran around 1500 lanes and 64% of libraries were sequenced in one lane, just 20% for >4 lanes. To get maximum efficiency from NovaSeq would require >80% of our sequencing to be run as “superpools” i.e. multiple users libraries pooled for each lane.
Only one sample can be run on each NovaSeq flowcell. It is unclear if we can safely pool libraries from different users, or whether there would be enough indexes available to maximize the efficiency gains on NovaSeq. We rely on users to submit libraries within certain guidelines but many fail to do so. This would be a much larger problem where libraries were pooled for sequencing as we may be using incorrect information for quantification.
A bigger problem is that pooling different library types is unlikely to be straightforward. Mixing RNA-Seq (average library size of 270bp +/-25bp), with a ChIP-Seq (average library size of 350bp +/-200bp) would be difficult. Mixing either of these with highly variable libraries like ATA-Seq, or a low-diversity and small insert-size libraries like smallRNA-Seq would increase the difficulties considerably.
The instrument implements an automatic wash after each run to reduce carry-over between runs.
It is unclear how comfortable users might be on mixing Chip, RNA, ATAC etc in a single pool as a super pool. Nor is it clear that Genologics LIMS can cope with multiple pools in a lane. This information will need to be collected and interpreted before any purchasing decisions can be made.
NovaSeq will reduce the costs of running larger RNA-Seq and exome projects,and would also allow small, medium, and possibly large, genome projects to be run cost and time efficiently in-house. The S4 flowcell should allow ~70-100 30-40x Human Genomes to be sequenced per week. This would reduce costs by about 20% since we currently pay VAT data delivered by service providers.
Illumina have mentioned that they are looking to increase the default number of barcodes available in their library prep kits from 96 to 384. However they have given no guarantees on timelines. This would reduce the need to pool different applications in a single run, although we would need to batch projects which could increase wait times for users. More indexes would have the biggest impact on the cost of RNA-seq, exome and sWGS projects.
2) Comparability between 4-colour and 2-colour SBS: as I pointed out in last weeks post there are still some issue with the 2-colour SBS that remain to be resolved. Illumina’s release of data is likely to be a big step towards reassuring users about the quality of NovaSeq data. However scientists are generally conservative in their approach to new technologies and I am sure many users will want to finish their experiments on HiSeq, rather than swap mid-project. But with projects getting larger and larger finding a convenient break can be tough!
Over time it looks like 2-colour SBS will become the dominant chemistry. So it is up to the community to get some of these comparisons performed and published. Hopefully the NIST team are already analysing the BaseSpace data from NA12878?
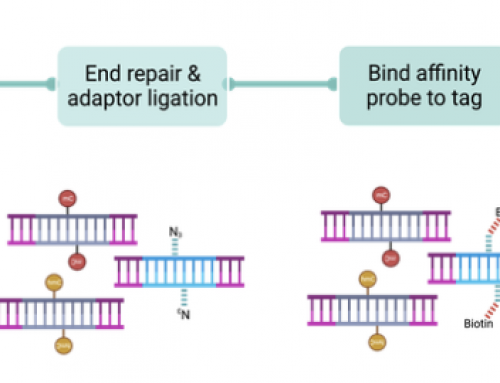
3) Fixing ExAmp clustering’s index swapping problems: Illumina’s Exclusion Amplification clustering chemistry was a big leap from the Manteia/Solexa developed bridge-amplification. It replaced the initial template annealing, followed by extension and cyclical amplification with a one-step isothermal process.
However ExAmp appears to have a problem – it results in swapping of indexes between samples in a multiplex pool. So reads from sample A in an A:B mix will contain a small percentage of reads from sample B and vice versa. This sounds terrible but is not a big problem for a standard RNA-seq project. But groups working on low-frequency allele detection should be understanding what impact this might have on their analysis.
This index-swapping is likely to be particularly bad for very deep multiplex pooled sequencing. In their customer facing slide deck Illumina suggest a 150,000x ctDNA genome, run as a single sample on an S4 flowcell. If more than one sample were pooled then index-swapping would be difficult to discern from low-frequency mutations in asymptomatic patients. I’ll assume sure Grail are working on this problem!
Perhaps the best fix for this problem is to use unique-at-both-ends dual-indexed libraries. In these the index mis-assignment at a low-level of around 1% will only results in 1% of 1% of samples being mis-assigned. Illumina have promised more indexes for NovaSeq to make RNA-Seq and Exomes even more affordable than they are today. I’m hoping these are provided as unique-at-both-ends dual-indexed adaptors.
Cost of experiments:
Most of the sequencing my lab runs is RNA-Seq, ChIP-Seq, Amplicomes and Exomes. The first two are counting applications where we use short single reads as the most cost-effective method to generate data, the most important consideration is often £per million reads. The other two require longer paired reads, here the most important consideration is often £per Gb.
RNA-Seq etc: We ran a very large number of “short-read” lanes in 2016. NovaSeq running S2 flowcells, each producing 3,300 million reads, results in 20% more reads per flowcell. Based on our 2016 sequencing requests this could result in 15% fewer flowcells being run. I suspect we’d run larger projects on NovaSeq so overall our reagent usage is likely to keep its upward trend.
However NovaSeq do not have a short-read S4 flowcell kit meaning users may generate paired-reads as standard. If there is no affect on costs then this is likely to be useful in identifying splices site and read duplicates more effectively than from single-end reads. The pro’s and con’s of single versus paired reads for RNA-Seq is a question that still elicits heated debate. If NovaSeq makes paired-end data the same cost as single-end that this might be the way the science goes. But an S4 run as long single-end might be even better for isoform detection.
Exomes etc: We ran a slightly smaller number of “long-read” lanes in 2016. NovaSeq running S2 flowcells, each producing 1Tb of data, results in a 23% price reduction in the cost per Gb. Again I suspect we’ll run more samples so Illumina won’t see the big drop in reagent volumes that accompanied the launch of the 600G sequencing chemistry.
According to Illumina there is a $2-4 per Gb difference ($16/18 vs $20) for NovaSeq S1/S2 flowcells versus HiSeq 4000. This is a reduction of 10-20% on current costs for consumables only. The price per million reads please is less clear as the costs of S1/S2 flowcell kits has not been released.
NovaSeq may be economical when service contracts are taken into consideration. A fleet of 2500’s could be replaced with 1 NovaSeq saving significant sums on those expensive HiSeq contracts. But this will reduce a pretty good revenue stream for Illumina as the numbers of NovaSeq in the field is unlikely to hit the highs of the 2500!
Overall costs are likely to drop for users as NovaSeq replaces HiSeq 2500 and 4000, and cannibalises HiSeq X Ten. But those costs apply primarily to consumables. As sequencing prices continue to plummet, library-prep costs get higher and higher (relatively speaking). Today library is at least 50% of the cost of doing RNA-Seq of ChIP-seq. In the future it might be as high as 75% of the cost. We desperately need library-prep innovation!
Mick Watson said on his NovaSeq post that these kind of figures mean a lot to large projects and large facilities. But for samller projects/facilities the need to immediately now is less clear.
Should your Genomics Core buy a NovaSeq?
The immediate answer is probably “no” since NovaSeq offers incremental gains over HiSeq 4000. However on the launch of the S4 flowcells we would make considerable savings against larger RNA-Seq and Exome projects. You’d also be able to run even large whole genome sequencing projects internally (and in the UK/EU this may save the costs of VAT). Of course the discount available to HiSeq 2500/4000 owners is likely to disappear by Jan 2018 so many labs will want to grab that saving.
With questions over 2-colour chemistry and especially the ongoing issue with index mis-assignment it remains to be seen whether people looking for rare variants in the clinical space will be won over by the new machine or not. I guess with Grail leading the way we should find out pretty soon!
This is such a big change in technology I suspect NovaSeq will be an instrument many core labs will look to purchase in 2018. And will very much depend on the kind of projects their users want to complete.















[…] the move to patterned flowcells and the issues the ExAmp clustering chemistry has (read this and this). If these turn out to be problems Illumina cannot resolve, or mitigate for customers, then the […]
Hi James,
I’m wondering what your take on the NovaSeq is now, after a year? It seems to me that NovaSeq is a replacement for a HiSeqX, but even with the XP Workflow, it’s not going to replace the HiSeq 4000 unless a new flowcell is released with 4 lanes, where each lane will cost the same or less than a HiSeq lane. (I’m told that a NovaSeq can only handle 4 lanes on 1 flow cell.)
Of your 3 NovaSeq limitations, at least #3 (index swapping) has been remedied.
#1, pooling, seems to still be the primary issue, even with the XP Workflow. A lane of PE 150bp on a NovaSeq costs more than double the S1 lane or quadruple the S2 or 4 flow cell lanes. Thus, the researcher must reorganize their workflow into much larger blocks of reads and $. As you pointed out, the problem for a core lab is that many researchers don’t need more than a lane of HiSeq 4000 and thus would have to index and pool multiple jobs into a lane. It doesn’t matter that the cost is less per base (the S1 is actually more per base with the XP Workflow) because you’re forced to buy more. It’s like shopping at a warehouse when all you need is a convenience store.
Unless Illumina makes a new flow cell, I think that many researchers will stick with HiSeq and thus even if Illumina stops making the HiSeq, it will continue to be used.
Has your take changed since january?
Cheers,
Chris
Hi Chris,
The additional costs are generally more than offset by the extra data the NovaSeq generates. However you are right that users will need to pile more samples into the lanes, or generate deeper coverage of the same number of samples. I think NovaSeq is the future for core labs – if they can afford it, and if their users require the increase in data. If a lab expected to create pools of multiple libraries to recognise the cost savings I’d expect them to run into big trouble. I doubt Illumina’s LIMS can handle the task, especially given that samples appear in the process already barcoded so you’d need to control what users did up front.
I’ll follow up with a post on RNA-Seq as I do think Illumina need to listen to customers about the inability to run a more cost effective single-end sequencing option. For counting applications we really do prefer more reads and single-end costs significantly less “per million reads” than paired-end.
I can’t edit my response.
replace the following sentence above:
A lane of PE 150bp on a NovaSeq costs more than double the S1 lane or quadruple the S2 or 4 flow cell lanes.
with this:
A lane of PE 150bp on a NovaSeq costs more than double the HiSeq4k (S1 lane is double and S4 lane is quadruple). Thus, the researcher must reorganize their workflow into much larger blocks of reads and $.
Hi James,
We have 50% of our RNA samples sequenced with HiSeq2500 and the other 50% with NovaSeq. Do we need to perform a particular normalization proocol in order to get those samples comparable?
Thanks
If the samples on 2500 will be analysed together, and the samples on 40000 will be analysed together, but the two data sets will not be directly analysed together then you’re probably OK. If you planned to analyse everything as one experiment in the first place then unfortunately you may have confounding factors.
Are your samples split across two groups? And if so is there one group per machine?