RNA-seq is getting very cool over in Sweden’s SciLifeLab. A recent press release announces that Joakim Lundeberg (Royal Institute of Technology) Jonas Frisén and Patrik StÃ¥hl (Karolinska Institutet) were awarded £1.8M from the Knut and Alice Wallenberg Foundation to study Brain biology. They have developed a method of 2-dimensional gene expression, 2D RNA-seq and will use this to comprehensively map gene expression in the brain. They also aim to compare normal and diseased brains, including cancer, autism and schizophrenia.
This method is likely to impact many other fields in genome biology. Being able to look and see how gene expression is localised in tissues allows us to move away from gross gene expression differences and understand how different cells and tissues interact with each other.
How does 2D-RNA-seq work: Patrik Ståhl presented a talk at AGBT, which I missed this year, fortunately Illumina,s Abizar Lakdawalla (Assoc Director, Novel DNA sequencing technologies and applications) blogged an overview.
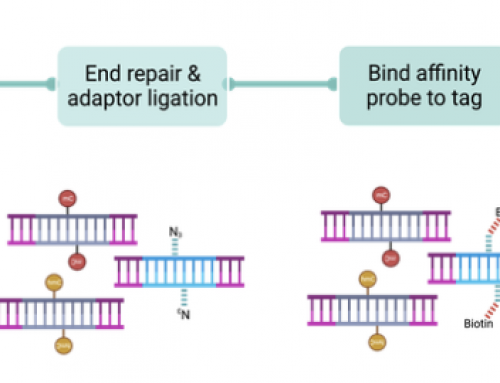
The method uses a surprisingly simple concept. A tissue section is cut and applied to a pretty standard microscope slide for histopathological analysis. However the slide is first prepared by spotting barcoded oligo-dT’s. These oligos capture RNA’s that diffuse directly onto the slide and are available for cDNA synthesis. The cDNAs can be processed to make an RNA-seq library, and the sequencing of the barcodes allows mapping of the reads back to the original slide image. Current slides contain over 100,000 images but there is no reason why higher density arrays can’t allow larger and more detailed sections to be studied.
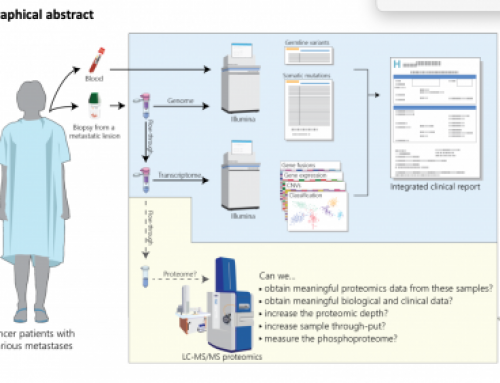
The image below was featured on Abizar’s blog, thanks to him for putting it out there!
 |
| 2D-RNA-seq |







great post! ill be keeping an eye on this blog.