This post has been very slightly modified from the 2012 original after getting a comment from a recent reader! Enjoy re-reading.
Eight years is a long time in NGS. I recently re-read a 2004 article in Pharmacogenomics 2004, and also found a EBI presentation from Clive Brown and Ewan Birney. Both of these were from a small company based in Cambridgeshire called Solexa. At the time of publication they had only just identified their first alpha-test site and the presentation talked about a prototype instrument ready for the end of 2004.
 |
| Prototpye GA1 |
Trademarks mentioned in the paper such as SMA-seq and TotalGenotyping have not, I suspect, been heard of by most Illumina sequencing users (including myself).
The paper describes where Solexa came from (Shankar Balasubramanian and David Klenerman’s patents of 1998 spun out of Cambridge University Department of Chemistry). It mentions Solexa’s demonstration version of “a system that will allow rapid, base-by-base comparison of genomic DNA sequences” and that this will produce “four or five orders of magnitude improvement over conventional sequencing”. Read lengths of just 25-30bp are proposed, and a nice graph illustrates how just over 80% of the Human genome is uniquely mappable with these incredibly short reads.
Simon Bennett, business development director of Solexa at the time and author of the Pharmacogenomics paper suggests that Solexa will achieve the $1000 genome within the next ten years. That leaves us two more years to get to $1000 genomes. It does not seem unreasonable that we’ll get there although more discussion today is about the cost of bioinformatics analysis!
What happened to single molecule sequencing: There is an overview of the Solexa Single Molecule ArrayTM technology that the paper suggest can analyse a Human genome in a single experiment. As described there were just 100,000 DNA molecules per cm2 compared to 100M per cm2 today. The basic chemistry description is unchanged from current SBS, although only 25 bases were being sequenced at the time of publication.
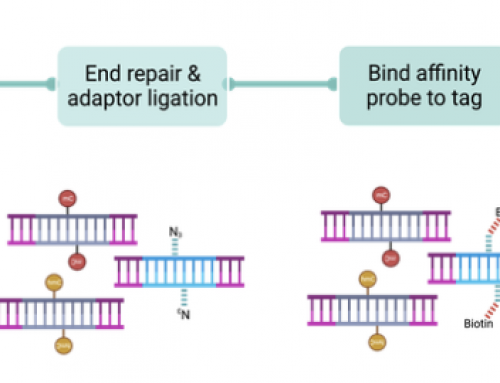
It is only towards the end of the paper that Solexa’s acquisition of Manteia’s solid surface bridge-amplification technology, this is the clustering we know and love today, and it gets slightly buried compared to Solexa’s invention “SmaSeq” – not the microbiome of baby formula, rather the acronym for Solexa chemistry prior to “SBS” . Up until this point Illumina had been focusing on single molecule sequencing. Without the acquisition of Manteia perhaps Solexa would have continued to chase single molecule sequencing and ended up like Helicos or Pacific BioSciences . As it stands clustering and SBS chemistry have been the bedrock of next-gen sequencing for the past five years.
Personally I’d bet Illumina are still putting lots of effort into single molecule approaches, and not just by investing in companies like ONT (seems a laugh now doesn’t it) . I’d like to know if it would be possible to sequence single molecules on a HiSeq with a more sensitive camera (massive oversimplification I know)? Imagine 1000M single molecule reads! This might not be what we ultimately use for single-molecule but I think we can be certain there is a lot more coming for next-gen in the next eight years.
PS: Would SOLiD have been the dominant technology if Agencourt had bought Manteia instead? Perhaps we should have a genomics version of Marvel’s “What if” comic books from the 80’s?
PPS: The Illumina history lesson also taught me that we share half our genes with bananas!








Hi James, it’s 2020. Any update on this? I am curious to see your views on SMS? Illumina’s take on SMS? ONT? and beyond?
Wow that was a blast from the past. I’ve updated the post slightly but I’d suggest you take a look at the announcements about CoolMPS from MGI. Read the coverage on GenomeWeb for a nice easy round-up.
Hi James, great post. I always enjoy looking back at the early days. At one point Illumina had SBS, SBL, ePCR and BridgePCR available. The stories how they came up with SBS/bridgePCR differ but my favorite is that they didn’t believe that SBL sequencing could be explained to customers. Did you know that the first “NGS” sequencer was actually from Lynx? MPSS..
https://www.ncbi.nlm.nih.gov/m/pubmed/10835600/
One of my early memories from Genomics was a talk by one of the Lynx founders. What they suggested seemed believable but impossible due to sequencing capacity. Oh to be blessed with better foresight!