A year ago I wrote a post about the explosion of different NGS acronyms. When I wrote it I was surprised to see over 30 different acronyms and suggested that part of this was authors wanting to make their work stand out, hopefully coining the next “ChIP-seqâ€.
In the past year more and more NGS acronyms have been published. I am partly responsible for one of these TAm-seq and understand better the reasoning for using acronyms. Once I have spoken to someone about the work we did in the STM paper I can simply refer to TAm-seq in future conversations.
It might help if we as a community could agree on a naming convention to make searching for work using specific techniques easier. There are multiple techniques for analysis of RNAs and using the catch-all “RNA-seq†would allow much quicker PubMed searching. Of course we would need to add keywords around the particular technique being used, RNA-seq could encompass mRNA, ribosome removal, strand-specific, small, micro, pi, linc, etc, etc, etc.
Here is a list of acronyms that we in the community could use to simplify things today. It would obviously need tidying up every year or so as new acronyms get added.
- DNA-seq: Unmodified genome sequencing.
- RNA-seq: All things RNA.
- SV-seq: Structural-variation sequencing.
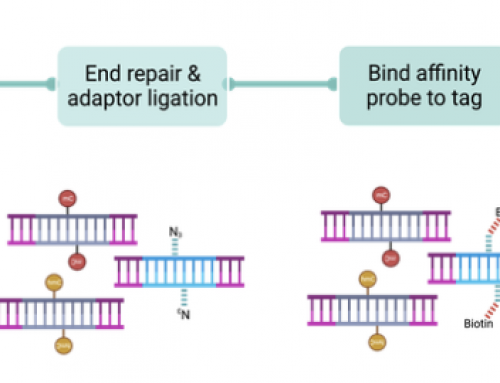
- Capture-seq: Exomes and other target capture sequencing.
- Amplicon-seq: Amplicon sequencing.
- Methyl-seq: Methylation and other base modification sequencing.
- IP-seq: Immuno-Precipitation sequencing.
Let me know what you think
Again, here is a link to the data.







I'm a newbie in NGS and am really confused what the difference between the Capture seq and the newer RNA capture seq, and also what differs between RNA Capture Seq and TruSeq (Multiplexing)? Any tips on what books to read?
Nora
a.thoeng@garvan.org.au