I’ve been finding out more about exomes: specifically QC analysis using HS Metrics in Picard. There’s loads of useful metrics and I’m hoping to get to a point that I can explain these to users here and also look at the results to try and troubleshoot an experiment. I’m also trying to understand what sort of read length we should be using for exome analysis. An earlier post discussed my thoughts around moving to PE125 or switching to SE125 and running more lanes. In a follow up post (watch this space) I’ll try to consider the impact of different run modes: will users/reviewers accept any kind of read for an exome or will they baulk at seeing something different from the paired-end norm?
PS: Your comments on this would be greatly appreciated!
The first thing I really wanted to get my head around was the difference between baits and targets. On the SAMtools mailing list many people suggest simply using the targets only, however that seems to defeat the purpose of having different files in the first place. Baits and targets are different; I’m guessing the companies that manufacture exome capture kits use an in-house algorithm to design baits from a list of targets and that differences in these may be somewhat immaterial if you only care about how well a single gene, e.g. TP53, is captured.
Baits and targets defined: A bait – is the biotinylated oligo-nucleotide or cRNA used to pull out fragments from a whole genome library. A target – is a region of the genome the manufacturer was trying to enrich for, it does not necessarily mean the capture will work a the target region, or that even if it does coverage will be even across different targets or samples!
Ideally we’d sequence the whole of a gene like TP53 as mutations can be found all over the place, however analysis of exons 4–10 covers well over 90% of mutations. The image below shows the genomic coordinates for exons 4-10 of TP53, and if you look closely you can just make out that baits are a little smaller than targets, and that some exons have multiple baits (exon 4).
 |
| click the image to navigate to the session. |
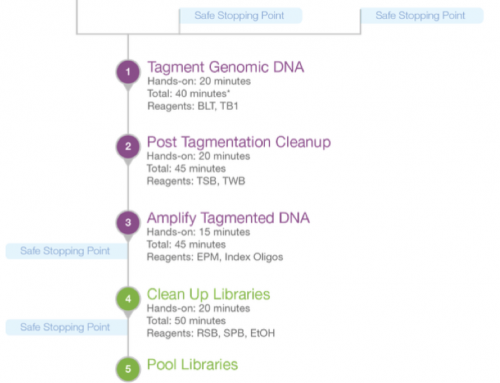
The distribution of target sizes is pretty high in the Nextera Rapid capture 1.2 we’re using in my lab. The total target size is 451Mb (using 336Mb of 79bp baits). The majority of targets are under 250bp, with almost 50% being under 150bp, or about the size of one exon. But 6% of targets are over 400bp and 10 are over 10Kb in length and very heavily tiled with baits.
HSmetrics: The tools in PICARD are really very useful, but did take a while for me to figure out. A 2010 presentation from the Broad’s Kristian Cibulskis and Andrew Kernytsky: Quality Assessment of Hybrid Selection Experiments, explains quite a lot about the Broad’s capture and analysis pipelines; and slides 9-12 cover interpretation of HS_metrics.
From a quality point of view a few of the metrics turn out to be very very useful (I’d appreciate some help filling in the numbers you think should be QC cutoffs!). Here are a few I think are most helpful…
PF_Unique_Reads [x%]: The number of PF reads that are not marked as duplicates
PF_UQ_Bases_Aligned [>90%]: the number of bases left after duplicate removal and alignment.
On_Target_Bases [x%]: the number of bases aligning to the targets file
PCT_Selected_Bases [x%]:On+Near Bait Bases / PF Bases Aligned.
Fold_enrichment [x%]: The fold by which the baited region has been amplified above genomic background.
A poster from AGBT ’13: Comparison of Illumina Sequencing Run Lengths and Genome References on Whole Exome Captured Libraries, by the Center for Inherited Disease Research (CIDR) in Baltimore shed some light on the move from PE75-100, and therefore helps me understand how the move to PE125 might impact us. They ran the same libraries on both run types and compared various exome analysis metrics. The PE100 data aligns better resulting in a slight increase in coverage.
I’ll be back with some comments on using different read-lengths for exomes: is there a “correct” choice?






Leave A Comment