Breast cancer researchers publish a lot of papers, and a large number of those will include some genomics, variant identification, RNA-Seq, ChIP-Seq, etc. Increasingly research is being carried out on patient derived material, but cell lines are still a vital part of much of the published research. Two things happened last year that are likely to have affected the breast cancer research community: Santa Cruz Biotech’s stopping antibody production, and the MCF-7 cell line being shown to have significant lot-to-lot, and even within-lot variation. These issues underscore the importance of replicating research, ideally using different reagents to see if biological effects are generally reproducible.
“Do scientists pay enough attention to the quality of their reagents?”
Antibody shutdown at Santa Cruz: Next year one of the most widely used antibodies in breast cancer research is pulled from the shelves. Santa Cruz Biotech’s ERα polyclonal antibody will no longer be available due to their license being revoked. The USDA issued a record fine of $3.5 million for alleged animal welfare violations, and Santa Cruz agreed to no longer sell antibodies made in USDA-regulated animal. Some antibodies were no longer produced from May of last year, and user will have been switching in the meantime. However Jan 2017 means anyone using this, and many other Santa Cruz antibodies, will be wholly reliant on their current stocks.
I’d not usually write about antibodies. However we see so much ChIP-seq coming through the lab I can see how important these reagents are to users. In experimental design discussions we often argue back-and-forth over the need to use multiple antibodies to the same target to reduce biases in experiments, or we’ve discussed the impact of using a different lot number in follow up experiments. The quality and continued supply of these antibodies is key to reproducible research efforts.
Santa Cruz Biotech’s ERα polyclonal antibody (HC-20 or sc-543) is widely used. A simple search on PubMed Central for papers using this particular antibody in a chromatin IP returns over 200 papers. Many of those research groups will have been working hard to find an alternative antibody, and to validate their earlier findings with this new reagent.
The Scientist published an antibody user satisfaction survey in 2012. This showed that Santa Cruz was number 2 in the Top 15 antibody providers (by sales), and that ChIP was number 8 in the Top 10 applications.
The survey did report on user satisfaction with the most highly rated companies generally being the smaller ones, Cell Signaling Technology, BioLegend, and eBioscience. Santa Cruz was rated as excellent by just 12% of users. It’s perhaps easy in hindsight but why were so many antibodies being ordered from a company so few people rated as excellent?
Whilst the study was published four years ago it did suggest that ChIP was the fastest growing application. I’d be interested to know what the list of top applications looks like today – Western blots are probably still number 1.
Heterogeneity of MCF-7: At about the same time as Santa Cruz were shutting down antibody production a paper was published in Nature’s Scientific Reports on the genetic variability in MCF-7 cells. MCF-7 is one of the most widely used breast cancer cell lines, being used for over 4 decades. It is an ER+ human breast adenocarcinoma cell line (actually a plural effusion) cited in over 20,000 articles. MCF-7, T-47D and MDA-MB-231 account for about two-thirds of abstracts. In the paper the authors showed significant heterogeneity in a single batch of cells, and importantly these cells had passed the standard STR cell authentication. Heterogeneity in MCF-7 has been previously reported, but many had assumed using cells from a verified source (e.g. ATCC), verifying them with STR profiling and keeping passage numbers low would remove most of the problems.

Figures B & C in the paper showed just how strikingly different two vials from the same ATCC lot were after passaging in two labs. There were significant differences in how the two sub-cultures proliferated after treatment with estrogen. Microarray, GX and CGH, and metabolomics analysis showed clear differences between the different cells. It is a shame that, given this work was published in 2016, the authors did not perform RNA-Seq, or run sWGS and/or exomes to determine SNVs and CNVs. Something is making cells from the same batch “tick” very differently and higher resolution genomic technologies may have been able to pinpoint what was different. Epigenetic effects were not discussed at all, but gene regulation could well have been affected.

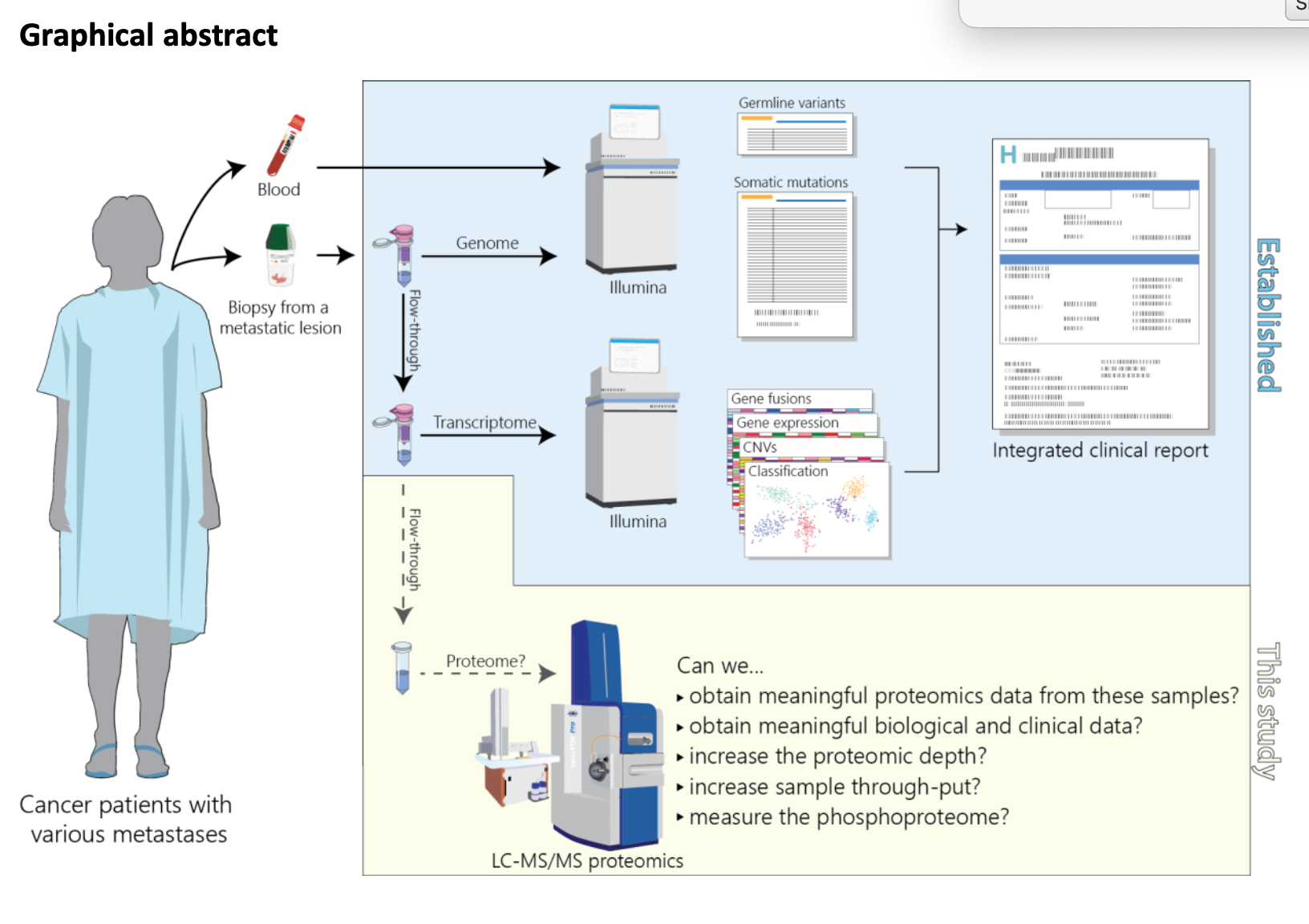




Surprised to still find novel transcripts in the MCF-7 cell line…
http://biorxiv.org/content/early/2017/01/17/100974
[…] factors than previously. Coupled to thefact that a specific antibody is not required (see this post describing just one issue with antibodies for ER, the most highly chipped factor in breast cancer research) means researchers can analyse TFs that […]