TIF-Seq
Transcript Isoform Sequencing
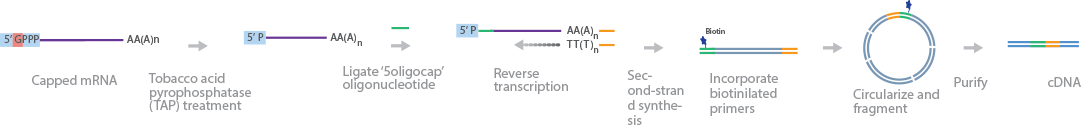
Transcript isoform sequencing (TIF-Seq) identifies transcript isoforms by selective sequencing of full-length mRNA molecules with 5’caps and poly(A) tails (Pelechano et al., 2013). Capped mRNA molecules are selected by substituting the 5’caps with oligonucleotides. To achieve this result, 5′-phosphate groups are removed from non-capped RNAs to differentiate them from their capped counterparts. The caps are removed by tobacco acid pyrophosphatase (TAP) treatment to expose the 5′-phosphate groups for ligation with oligonucleotides. mRNAs are separated into 2 different tubes and reverse-transcribed to generate full-length cDNA (flcDNA). flcDNA in each tube is annealed to barcoded 5′-biotinylated primers and 3′ primers. The primers contain barcode sequences unique to each tube, as a control mechanism against chimeric fragments and intermolecular ligation. The 2 tubes are combined, digested with NotI enzyme to produce sticky ends, and ligated to form circular double-stranded cDNA, which is fragmented subsequently. Fragments containing the biotinylated 3′ and 5′ ends are isolated with streptavidin. Multiplexing barcodes are added to the 3′ and 5′ ends of the purified cDNA fragments to create a cDNA library for sequencing.
Advantages:
- Identifies transcript isoforms by sequencing the 5′ and 3′ end of the same RNA strand
- Chimera-control barcodes filter out intermolecular ligation of cDNA fragments
Disadvantages:
- Requires a large amount of full-length RNA in the sample
- Use of NotI enzyme to introduce sticky ends is only viable for AT-rich genomes like yeast
- Strong bias toward short RNA strands (Pelechano et al., 2014)
Reagents:
Illumina Library prep and Array Kit Selector
Reviews:
Anamika K., Verma S., Jere A. and Desai A. Transcriptomic Profiling Using Next Generation Sequencing – Advances, Advantages, and Challenges. 2016;
Bagchi D. N. and Iyer V. R. The Determinants of Directionality in Transcriptional Initiation. Trends Genet. 2016;32:322-333
Sole C., Nadal-Ribelles M., de Nadal E. and Posas F. A novel role for lncRNAs in cell cycle control during stress adaptation. Curr Genet. 2015;61:299-308
References:
Pelechano V., Wei W. and Steinmetz L. M. Extensive transcriptional heterogeneity revealed by isoform profiling. Nature. 2013;497:127-131
Pelechano V., Wei W., Jakob P. and Steinmetz L. M. Genome-wide identification of transcript start and end sites by transcript isoform sequencing. Nat Protoc. 2014;9:1740-1759
Related
History: TIF-Seq
Revision by sbrumpton on 2017-06-21 07:50:24 - Show/Hide
Transcript Isoform Sequencing

Transcript isoform sequencing (TIF-Seq) identifies transcript isoforms by selective sequencing of full-length mRNA molecules with 5'caps and poly(A) tails (Pelechano et al., 2013). Capped mRNA molecules are selected by substituting the 5'caps with oligonucleotides. To achieve this result, 5'-phosphate groups are removed from non-capped RNAs to differentiate them from their capped counterparts. The caps are removed by tobacco acid pyrophosphatase (TAP) treatment to expose the 5'-phosphate groups for ligation with oligonucleotides. mRNAs are separated into 2 different tubes and reverse-transcribed to generate full-length cDNA (flcDNA). flcDNA in each tube is annealed to barcoded 5'-biotinylated primers and 3' primers. The primers contain barcode sequences unique to each tube, as a control mechanism against chimeric fragments and intermolecular ligation. The 2 tubes are combined, digested with NotI enzyme to produce sticky ends, and ligated to form circular double-stranded cDNA, which is fragmented subsequently. Fragments containing the biotinylated 3' and 5' ends are isolated with streptavidin. Multiplexing barcodes are added to the 3' and 5' ends of the purified cDNA fragments to create a cDNA library for sequencing.
Advantages:- Identifies transcript isoforms by sequencing the 5' and 3' end of the same RNA strand
- Chimera-control barcodes filter out intermolecular ligation of cDNA fragments
Disadvantages:- Requires a large amount of full-length RNA in the sample
- Use of NotI enzyme to introduce sticky ends is only viable for AT-rich genomes like yeast
- Strong bias toward short RNA strands (Pelechano et al., 2014)
Reagents:Illumina Library prep and Array Kit SelectorReviews:Anamika K., Verma S., Jere A. and Desai A. Transcriptomic Profiling Using Next Generation Sequencing - Advances, Advantages, and Challenges. 2016;Bagchi D. N. and Iyer V. R. The Determinants of Directionality in Transcriptional Initiation. Trends Genet. 2016;32:322-333Sole C., Nadal-Ribelles M., de Nadal E. and Posas F. A novel role for lncRNAs in cell cycle control during stress adaptation. Curr Genet. 2015;61:299-308References:Pelechano V., Wei W. and Steinmetz L. M. Extensive transcriptional heterogeneity revealed by isoform profiling. Nature. 2013;497:127-131Pelechano V., Wei W., Jakob P. and Steinmetz L. M. Genome-wide identification of transcript start and end sites by transcript isoform sequencing. Nat Protoc. 2014;9:1740-1759
