SHAPE-MaP
Selective 2ê-Hydroxyl Acylation Analyzed by Primer Extension and Mutational Profiling
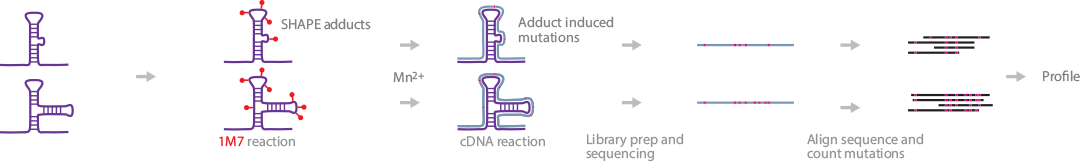
SHAPE-MaP sequences secondary RNA structures at various levels on a massively parallel scale. The method can be customized to interrogate small RNAs, amplicons, or rare RNA species accurately in a mixture of RNAs (Siegfried et al., 2014) (Smola et al., 2015). As implied in the name, SHAPE-Map uses the SHAPE-Seq 1M7 reaction to mark RNA ribose 2ê-OH groups to identify secondary RNA structures. This reaction is followed by mutational profiling (MaP) to induce noncomplementary nucleotide mutations on SHAPE adducts during RT. These MaP mutations are analyzed after sequencing using powerful informatics tools developed specifically for this method.
Briefly, SHAPE electrophiles are added to the sample containing folded RNAs. The samples are divided into 3 different reaction lines: +reagent, _reagent, and a denaturing control, to correct against intrinsic background mutation rates from reverse transcription. After SHAPE mutations are introduced, RT primers are selected depending on the RNA type of interest (amplicon, small RNA, or specific RNA species profiling). The primers are added to the reaction and the RNA is reverse-transcribed. Subsequent library preparation steps differ according to the RNA of interest. The small RNA workflow involves standard PCR amplification with appropriate primers. The workflows for amplicons and specific RNA species follow the Nextera XT DNA Library Preparation Kit protocol. The barcoded samples are ready for sequencing.
Similar methods: SHAPE-Seq, icSHAPE
Advantages:
- Identifies RNA structures at single-nucleotide resolution using a combination of SHAPE-Seq and MaP techniques
- Library preparation step is highly customizable for different applications: amplicon, small RNA, or rare RNA species profiling
- Avoids common issues encountered with adapter-ligation methods
Disadvantages:
- Length of the RNA strand must be at least ~150 nt for the randomer and native workflow, and at least ~40 nt for the small-RNA workflow
- Difficult to distinguish MaP mutations from background noise in samples with low levels of RNA
Reagents:
Illumina Library prep and Array Kit Selector
Reviews:
Schmitz S. U., Grote P. and Herrmann B. G. Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci. 2016;73:2491-2509
Kwok C. K. Dawn of the in vivo RNA structurome and interactome. Biochem Soc Trans. 2016;44:1395-1410
Smola M. J., Calabrese J. M. and Weeks K. M. Detection of RNA-Protein Interactions in Living Cells with SHAPE. Biochemistry. 2015;54:6867-6875
References:
Mauger D. M., Golden M., Yamane D., et al. Functionally conserved architecture of hepatitis C virus RNA genomes. Proc Natl Acad Sci U S A. 2015;112:3692-3697
Lu Y. F., Mauger D. M., Goldstein D. B., Urban T. J., Weeks K. M. and Bradrick S. S. IFNL3 mRNA structure is remodeled by a functional non-coding polymorphism associated with hepatitis C virus clearance. Sci Rep. 2015;5:16037
Taylor A. I., Pinheiro V. B., Smola M. J., et al. Catalysts from synthetic genetic polymers. Nature. 2014;
Related
History: SHAPE-MaP
Revision by sbrumpton on 2017-06-21 09:06:28 - Show/Hide
Selective 2ê-Hydroxyl Acylation Analyzed by Primer Extension and Mutational Profiling

SHAPE-MaP sequences secondary RNA structures at various levels on a massively parallel scale. The method can be customized to interrogate small RNAs, amplicons, or rare RNA species accurately in a mixture of RNAs (Siegfried et al., 2014) (Smola et al., 2015). As implied in the name, SHAPE-Map uses the SHAPE-Seq 1M7 reaction to mark RNA ribose 2ê-OH groups to identify secondary RNA structures. This reaction is followed by mutational profiling (MaP) to induce noncomplementary nucleotide mutations on SHAPE adducts during RT. These MaP mutations are analyzed after sequencing using powerful informatics tools developed specifically for this method.
Briefly, SHAPE electrophiles are added to the sample containing folded RNAs. The samples are divided into 3 different reaction lines: +reagent, _reagent, and a denaturing control, to correct against intrinsic background mutation rates from reverse transcription. After SHAPE mutations are introduced, RT primers are selected depending on the RNA type of interest (amplicon, small RNA, or specific RNA species profiling). The primers are added to the reaction and the RNA is reverse-transcribed. Subsequent library preparation steps differ according to the RNA of interest. The small RNA workflow involves standard PCR amplification with appropriate primers. The workflows for amplicons and specific RNA species follow the Nextera XT DNA Library Preparation Kit protocol. The barcoded samples are ready for sequencing.
Similar methods: SHAPE-Seq, icSHAPE
Advantages:- Identifies RNA structures at single-nucleotide resolution using a combination of SHAPE-Seq and MaP techniques
- Library preparation step is highly customizable for different applications: amplicon, small RNA, or rare RNA species profiling
- Avoids common issues encountered with adapter-ligation methods
Disadvantages:- Length of the RNA strand must be at least ~150 nt for the randomer and native workflow, and at least ~40 nt for the small-RNA workflow
- Difficult to distinguish MaP mutations from background noise in samples with low levels of RNA
Reagents:Illumina Library prep and Array Kit SelectorReviews:Schmitz S. U., Grote P. and Herrmann B. G. Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci. 2016;73:2491-2509Kwok C. K. Dawn of the in vivo RNA structurome and interactome. Biochem Soc Trans. 2016;44:1395-1410Smola M. J., Calabrese J. M. and Weeks K. M. Detection of RNA-Protein Interactions in Living Cells with SHAPE. Biochemistry. 2015;54:6867-6875References:Mauger D. M., Golden M., Yamane D., et al. Functionally conserved architecture of hepatitis C virus RNA genomes. Proc Natl Acad Sci U S A. 2015;112:3692-3697Lu Y. F., Mauger D. M., Goldstein D. B., Urban T. J., Weeks K. M. and Bradrick S. S. IFNL3 mRNA structure is remodeled by a functional non-coding polymorphism associated with hepatitis C virus clearance. Sci Rep. 2015;5:16037Taylor A. I., Pinheiro V. B., Smola M. J., et al. Catalysts from synthetic genetic polymers. Nature. 2014;
