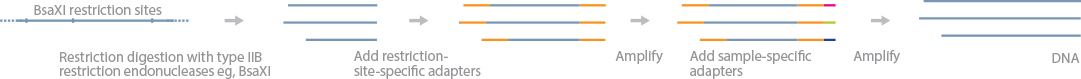2b-RAD
RAD With Type IIB Restriction Endonucleases
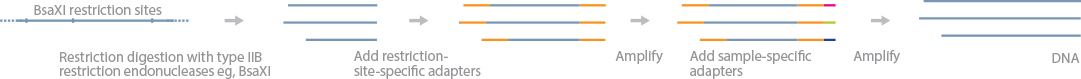
2b-RAD is similar to ddRadseq but uses type IIB restriction enzymes (BsaXI or AlfI), which will cleave upstream and downstream of a recognition site(Wang et al., 2012) . This shears the target genome into a large number of DNA fragments with a constant length of 33 bp (BsaXI) or 36 bp (AlfI). These short DNA fragments can be sequenced to determine genetic variants.
In this method, gDNA is first digested with a restriction enzyme (BsaXI), and adapters with partial (NNN) overhangs are ligated to the fragments. The adapter-ligated fragments from different samples are combined, and the fragments are amplified to produce the sequencing library.
Advantages:
- Highly reduced 2b-RAD libraries require much less sequencing for accurate genotyping
- High density of markers
- No interim purification steps, reducing losses and processing time
Disadvantages:
- Requires a reference genome
- Short tags may not be long enough for efficient locus discrimination in complex genomes
Reagents:
Illumina Library prep and Array Kit Selector
Reviews:
Andrews K. R., Good J. M., Miller M. R., Luikart G. and Hohenlohe P. A. Harnessing the power of RADseq for ecological and evolutionary genomics. Nat Rev Genet. 2016;
da Fonseca R. R., Albrechtsen A., Themudo G. E., Ramos-Madrigal J., Sibbesen J. A., et al. Next-generation biology: Sequencing and data analysis approaches for non-model organisms. Mar Genomics. 2016;30:3-13
Jiang N., Zhang F., Wu J., Chen Y., Hu X., et al. A highly robust and optimized sequence-based approach for genetic polymorphism discovery and genotyping in large plant populations. Theor Appl Genet. 2016;129:1739-1757
Kagale S., Koh C., Clarke W. E., Bollina V., Parkin I. A., et al. Analysis of Genotyping-by-Sequencing (GBS) Data. Methods Mol Biol. 2016;1374:269-284
Manel S., Perrier C., Pratlong M., Abi-Rached L., Paganini J., et al. Genomic resources and their influence on the detection of the signal of positive selection in genome scans. Mol Ecol. 2016;25:170-184
References:
Pecoraro C., Babbucci M., Villamor A., et al. Methodological assessment of 2b-RAD genotyping technique for population structure inferences in yellowfin tuna (Thunnus albacares). Mar Genomics. 2016;25:43-48
Fu B., Liu H., Yu X. and Tong J. A high-density genetic map and growth related QTL mapping in bighead carp (Hypophthalmichthys nobilis). Sci Rep. 2016;6:28679
Related
History: 2b-RAD
Revision by sbrumpton on 2017-06-21 07:50:21 - Show/Hide
RAD With Type IIB Restriction Endonucleases
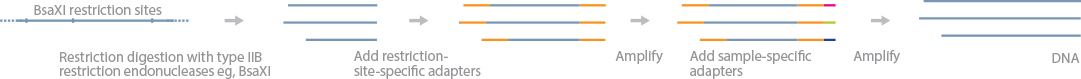
2b-RAD is similar to ddRadseq but uses type IIB restriction enzymes (BsaXI or AlfI), which will cleave upstream and downstream of a recognition site(Wang et al., 2012) . This shears the target genome into a large number of DNA fragments with a constant length of 33 bp (BsaXI) or 36 bp (AlfI). These short DNA fragments can be sequenced to determine genetic variants.
In this method, gDNA is first digested with a restriction enzyme (BsaXI), and adapters with partial (NNN) overhangs are ligated to the fragments. The adapter-ligated fragments from different samples are combined, and the fragments are amplified to produce the sequencing library.
Advantages:- Highly reduced 2b-RAD libraries require much less sequencing for accurate genotyping
- High density of markers
- No interim purification steps, reducing losses and processing time
Disadvantages:- Requires a reference genome
- Short tags may not be long enough for efficient locus discrimination in complex genomes
Reagents:Illumina Library prep and Array Kit SelectorReviews:Andrews K. R., Good J. M., Miller M. R., Luikart G. and Hohenlohe P. A. Harnessing the power of RADseq for ecological and evolutionary genomics. Nat Rev Genet. 2016;da Fonseca R. R., Albrechtsen A., Themudo G. E., Ramos-Madrigal J., Sibbesen J. A., et al. Next-generation biology: Sequencing and data analysis approaches for non-model organisms. Mar Genomics. 2016;30:3-13Jiang N., Zhang F., Wu J., Chen Y., Hu X., et al. A highly robust and optimized sequence-based approach for genetic polymorphism discovery and genotyping in large plant populations. Theor Appl Genet. 2016;129:1739-1757Kagale S., Koh C., Clarke W. E., Bollina V., Parkin I. A., et al. Analysis of Genotyping-by-Sequencing (GBS) Data. Methods Mol Biol. 2016;1374:269-284Manel S., Perrier C., Pratlong M., Abi-Rached L., Paganini J., et al. Genomic resources and their influence on the detection of the signal of positive selection in genome scans. Mol Ecol. 2016;25:170-184References:Pecoraro C., Babbucci M., Villamor A., et al. Methodological assessment of 2b-RAD genotyping technique for population structure inferences in yellowfin tuna (Thunnus albacares). Mar Genomics. 2016;25:43-48Fu B., Liu H., Yu X. and Tong J. A high-density genetic map and growth related QTL mapping in bighead carp (Hypophthalmichthys nobilis). Sci Rep. 2016;6:28679